New publication in "Nucleic Acids Research"
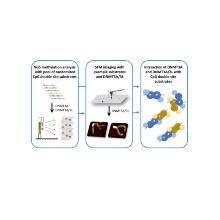
Heterotetramers of the DNMT3A/3L DNA methyltransferase contain two active centers binding CpG target sites at 12 bp distance, however their interaction with DNA not containing this feature is unclear. In this publication, a kinetic and SFM study of the Jeltsch group and the group of Dr. Tessmer at University of Würzburg was conducted. with this combined approach, we identified novel dimeric arrangements of DNMT3A or DNMT3A/3L tetramers on the DNA in a side-by-side and tetramer-swap mode. This variability in complex quaternary structures leads to a high adaptability in the interactions of DNMT3A and DNMT3A/3L with DNA, with preferential co-methylation not only of CpG sites at distances of ~12-13 bp, but also at ~2-3, ~5-6, or ~8-9 bp. These flexible modes of DNA interactions explain how DNMT3A and DNMT3A/3L can overcome their in-built structural preference for interaction with CpG sites with 12 bp spacing and introduce methylation into cellular DNA efficiently.