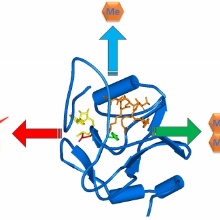
Like other important regulatory proteins, Protein Lysine Methyltransferases are subject to somatic mutations in tumors. The MLL3 enzyme monomethylates Lysine 4 in Histone H3. It is a frequently mutated enzyme, but the effects of cancer mutations in this protein were not yet studied. We show here that mutations found in the catalytic SET domain of MLL3 have defined and converse effects on MLL3. The N4848S mutation leads to a loss of the catalytic activity of MLL3, which resembles the effect of other loss of function mutation in MLL3 protein, like frameshifts or loss of expression. Such mutations may lead to the loss of H3K4 methylation at target genes and inhibit the expression of tumor suppressor genes. The Y4884C mutation leads to a change in the substrate specificity and product pattern of MLL3. This may result in the deposition of aberrant H3K4 trimethylation at enhancers leading to their conversion to promoters and the expression of oncogenes. Our data uncover potential approaches for targeted therapy, e.g. the application of an MLL3 inhibitor in patients carrying the Y4884C mutation. On a more general scale our data illustrate that cancer mutations need to be functionally studied in order to define their effects and based on this develop and apply individual treatments.