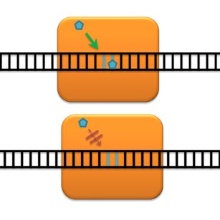
DNA methylation patterns are essential epigenetic Signals, which are inherited through cell division. The Dnmt1 DNA Methyltransfease plays a central role in this process. The DFG has decided on June 19th 2012 to support our investigation of the properties and mechanism of this enzyme over the next three years. In this project, we address the question that the in vitro activity of Dnmt1 is not sufficient to explain the speed of remethylation of DNA after DNA replication in cells. Preliminary data from our lab suggest that the in vitro rate of Dnmt1 is underestimated, because the enzyme exists in different conformations. Novel structural data indeed show that Dnmt1 can adopt different autoinhibitory conformations, in which other domains block access of the DNA to the catalytic domain. We plan to carry out single enzyme kinetics and determine the DNA methylation activity of Dnmt1 in its most active conformation after DNA has been bound. In addition, we aim to measure if a possible direct movement of Dnmt1 along DNA in cells (as opposed to the random walk on DNA in vitro) could further speed up DNA methylation. The starting point of the second part of our proposal is that the in vitro specificity of Dnmt1 is obviously not sufficient to ensure correct copying of methylation patterns without additional regulation, because the enzyme also shows activity towards unmethylated CpG sites. This is particularly critical outside of S-phase, when no “ correct” hemimethylated CpG substrate sites are available. Therefore, we aim to study if cell cycle coupled regulation of Dnmt1 occurs. To this end, we plan to identify post-translational modifications of Dnmt1 and study their cell cycle connection. We will further study, whether they influence the enzyme’s activity or specificity or its interaction with other protein.
Recent publications: